Introduction
I decided to try Neptune, AWS’s fully managed graph database service. The plan is to go from instance creation through loading RDF-format data, and finally issue a simple query using SPARQL.
I’ll cover what a graph database is and what Amazon Neptune is in a separate article.
Steps to be performed:
- Create the instance
- Create an IAM role, attach the role to Neptune, configure the S3 VPC endpoint
- Load data from S3
- Verify the loaded data using the RDF4J console and HTTP REST endpoint
Prerequisites:
- VPC and S3 created in advance
Creating the Instance
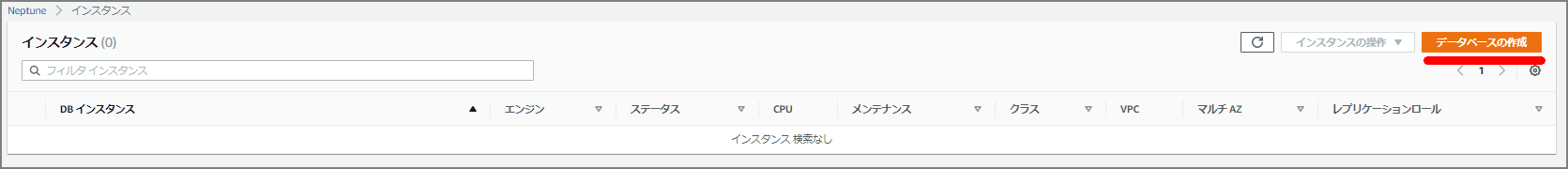
Select “Create Database”
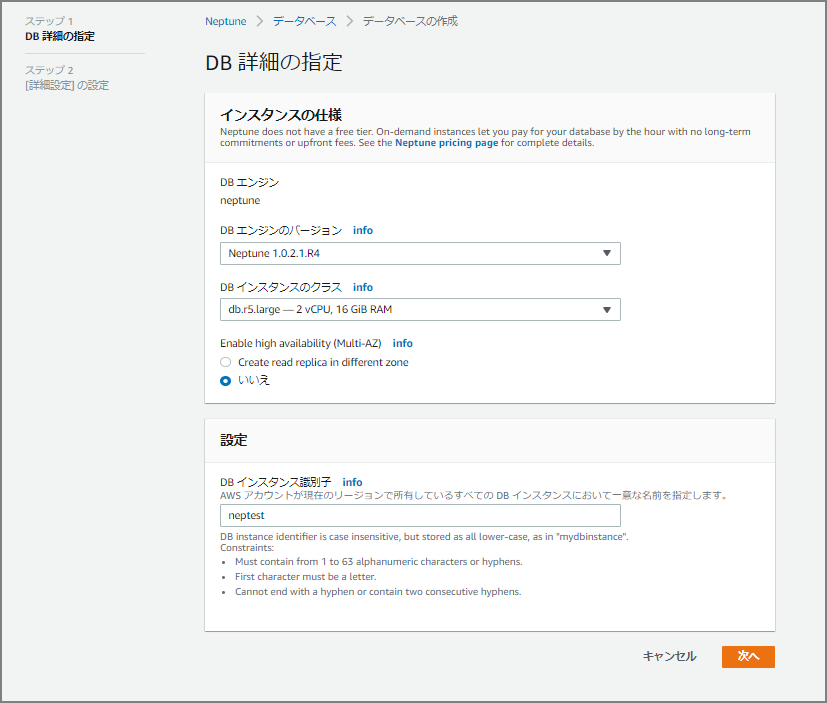
Fill in “Specify DB Details”
For this test, I specified “Neptune 1.0.2.1.R4,” the latest version at the time. Note that changing to Multi-AZ after instance creation is not currently possible, so select it at this stage if needed.
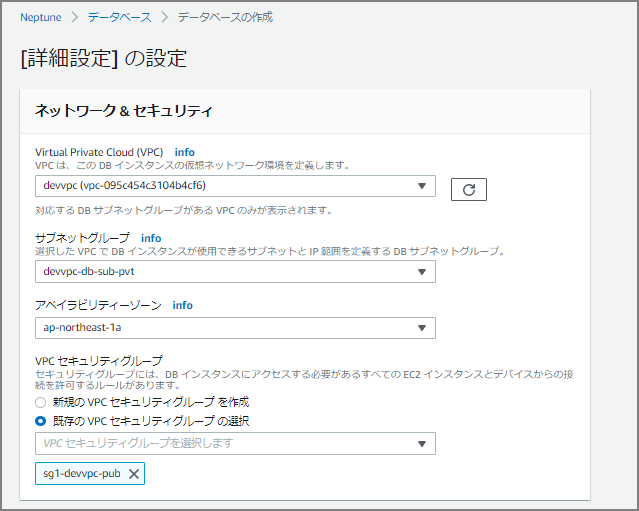
Continue Filling In Details
The input fields are similar to RDS and Aurora.
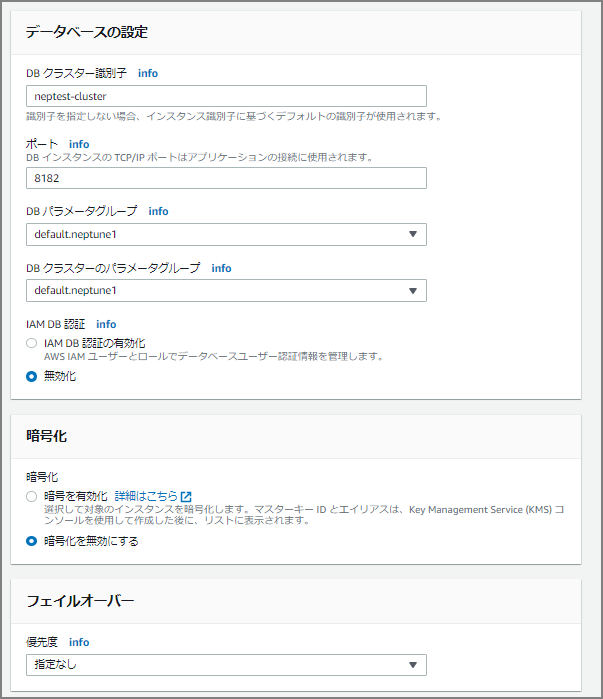
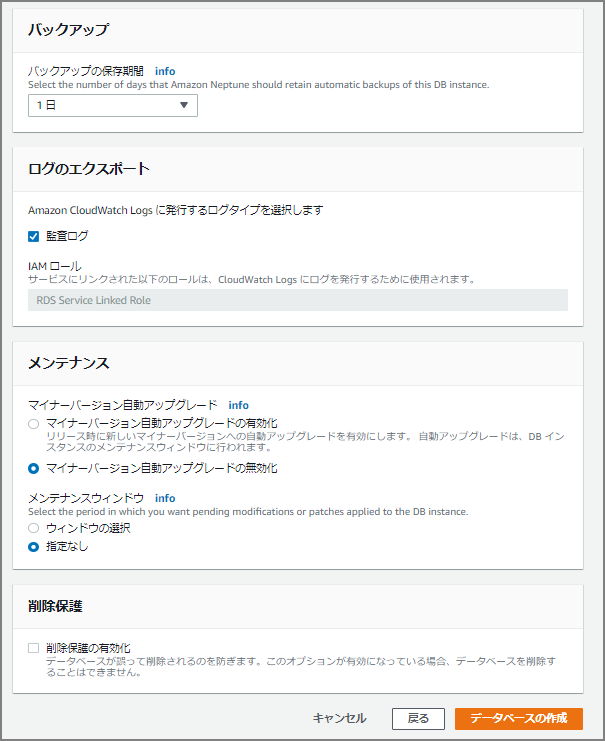
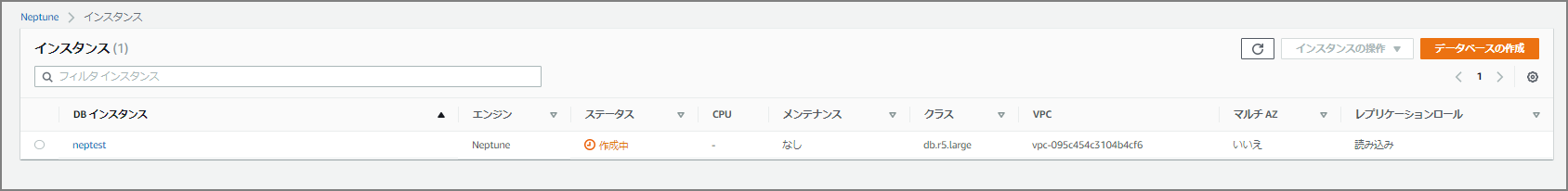
After clicking the “Create Database” button, creation begins — wait a bit.
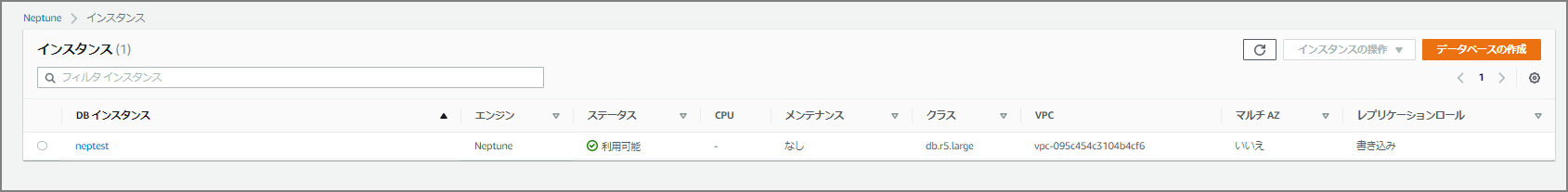
Creation completed in roughly 5 to 10 minutes.
Setting Up IAM Role and S3 VPC Endpoint
As preparation for data loading, configure the IAM role and S3 VPC endpoint.
Prerequisite: IAM Role and Amazon S3 Access - Amazon Neptune https://docs.aws.amazon.com/ja_jp/neptune/latest/userguide/bulk-load-tutorial-IAM.html
From the IAM screen, select “Create Role.”
Select S3.
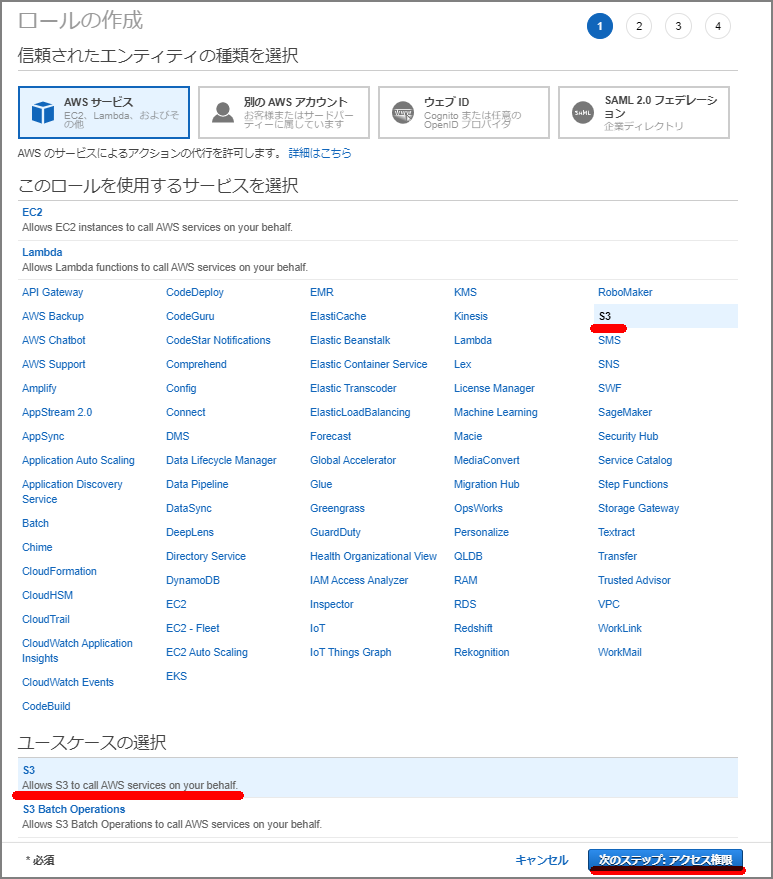
Select “AmazonS3ReadOnlyAccess” to attach the policy.
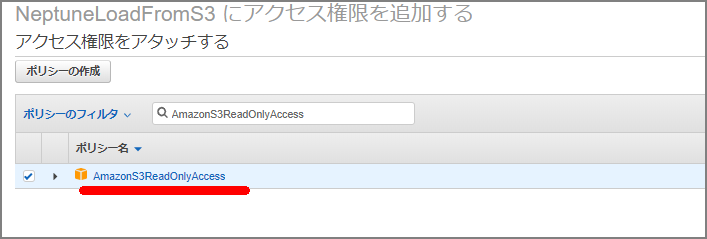
Fill in as needed.
The role name was set to “NeptuneLoadFromS3.”
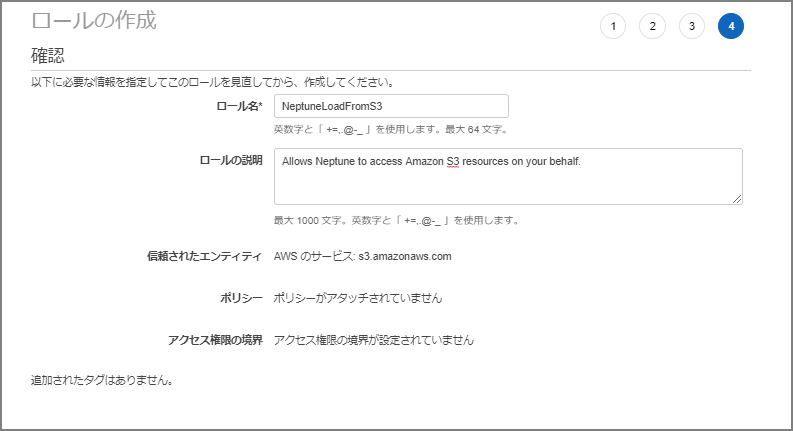
Navigate to the created role’s screen.
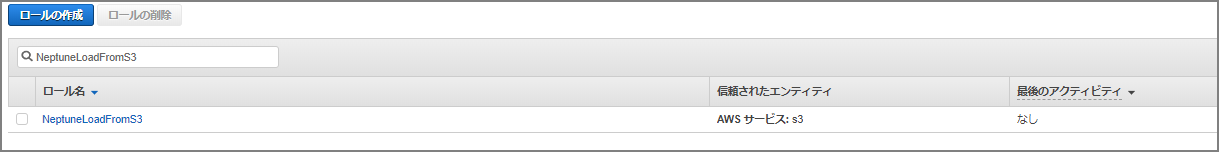
Go to “Trust relationships” - “Edit trust relationship” and paste the following, overwriting the existing content.
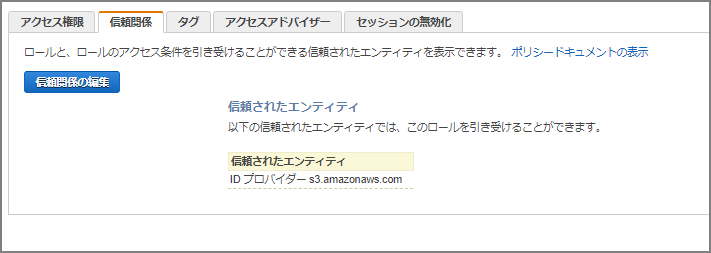
{
"Version": "2012-10-17",
"Statement": [
{
"Sid": "",
"Effect": "Allow",
"Principal": {
"Service": [
"rds.amazonaws.com"
]
},
"Action": "sts:AssumeRole"
}
]
}
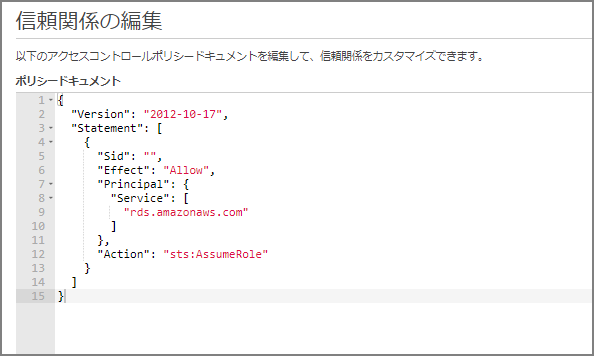
Add an IAM Role to the Amazon Neptune Cluster
Go to the Neptune cluster and select “Manage IAM roles.”
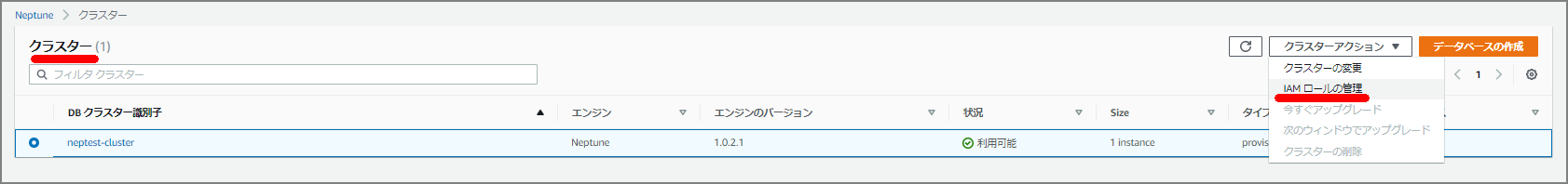
Add the IAM role just created (NeptuneLoadFromS3).
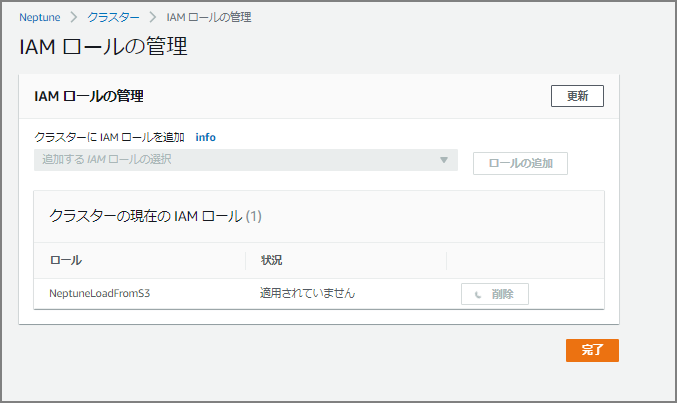
Create an S3 VPC Endpoint
A VPC endpoint is required to load data from S3 into Neptune. Set up a VPC endpoint.
On the endpoint creation screen, select “com.amazonaws.ap-northeast-1.s3.” (Since this is the Tokyo region, it’s ap-northeast-1; other regions will have a different region name.)
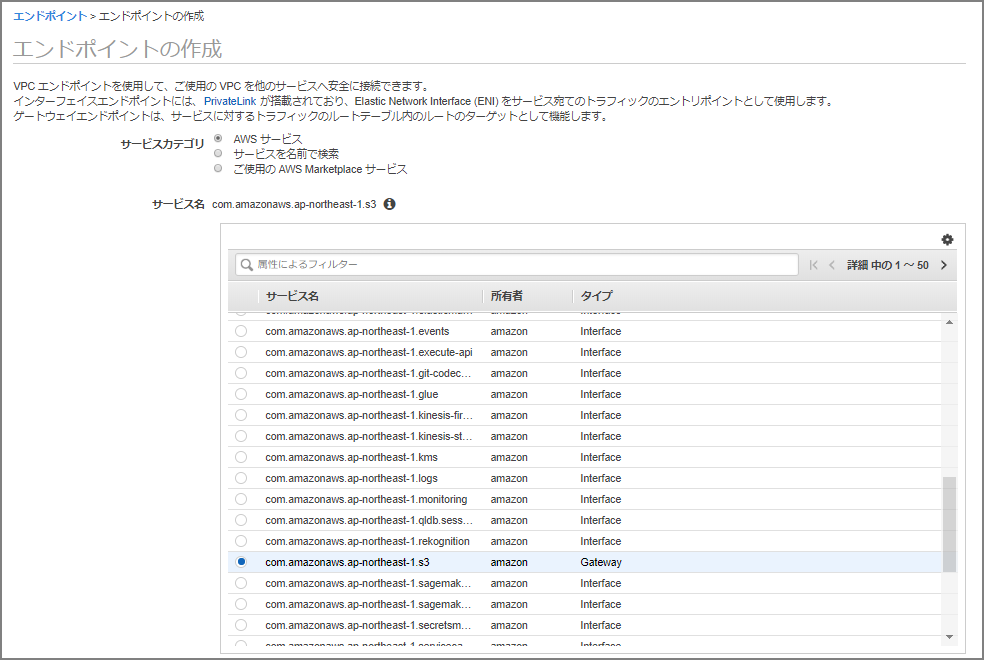
Specify the VPC and route table.
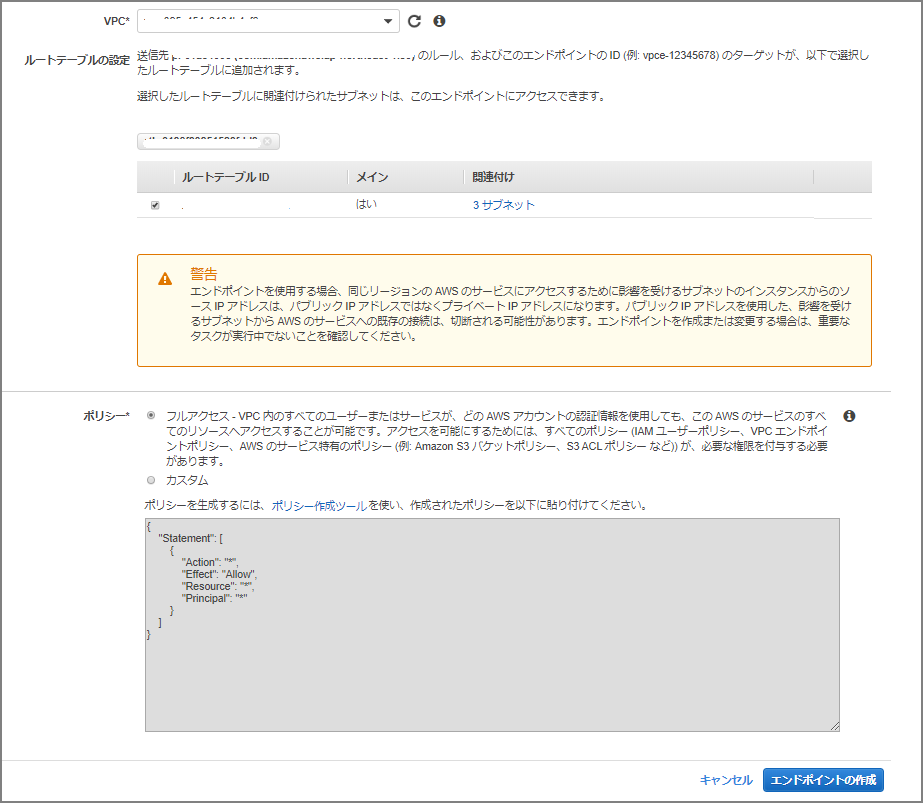
Loading Data from S3 into Neptune
We’re now ready to load from S3 into Neptune. The data to load will be from http://rdf.geospecies.org . Upload a sample RDF data file in rdfxml format to the designated S3 bucket.
[ec2-user@bastin nep-tool]$ curl -O http://rdf.geospecies.org/geospecies.rdf.gz
% Total % Received % Xferd Average Speed Time Time Time Current
Dload Upload Total Spent Left Speed
0 0 0 0 0 0 0 0 --:--:-- --:--:-- --:--:-- 0
100 8891k 100 8891k 0 0 3405k 0 0:00:02 0:00:02 --:--:-- 3404k
[ec2-user@bastin nep-tool]$
[ec2-user@bastin nep-tool]$ ls -l geospecies.rdf.gz
-rw-rw-r-- 1 ec2-user ec2-user 9105109 Jan 28 08:16 geospecies.rdf.gz
[ec2-user@bastin nep-tool]$ aws s3 cp geospecies.rdf.gz s3://nep-s3-xxxx/
upload: ./geospecies.rdf.gz to s3://nep-s3-xxxx/geospecies.rdf.gz
[ec2-user@bastin nep-tool]$
Load the data with the following command. Update endpoint, source, format, and iamRoleArn as needed.
For RDF, the format can also be turtle, ntriples, and others.
Load Data Formats - Amazon Neptune https://docs.aws.amazon.com/ja_jp/neptune/latest/userguide/bulk-load-tutorial-format.html
curl -X POST \
-H 'Content-Type: application/json' \
https://neptest.xxxxxxxxxxxx.ap-northeast-1.neptune.amazonaws.com:8182/loader -d '
{
"source" : "s3://nep-s3-xxxx/geospecies.rdf.gz",
"format" : "rdfxml",
"iamRoleArn" : "arn:aws:iam::xxxxxxxxx:role/NeptuneLoadFromS3",
"region" : "ap-northeast-1",
"failOnError" : "FALSE",
"parallelism" : "HIGH"
}'
After execution, the following is displayed. Note the loadId as it’s needed to check status.
{
"status" : "200 OK",
"payload" : {
"loadId" : "eff1268f-17ab-473a-b845-c2d91a317c01"
}
Check the data load status. Specify the loadId obtained earlier.
curl -G 'https://neptest.xxxxxxxxxxxx.ap-northeast-1.neptune.amazonaws.com:8182/loader/eff1268f-17ab-473a-b845-c2d91a317c01'
In-Progress Output
[ec2-user@bastin nep-tool]$ curl -G 'https://neptest.xxxxxxxxxxxx.ap-northeast-1.neptune.amazonaws.com:8182/loader/eff1268f-17ab-473a-b845-c2d91a317c01'
{
"status" : "200 OK",
"payload" : {
"feedCount" : [
{
"LOAD_IN_PROGRESS" : 1
}
],
"overallStatus" : {
"fullUri" : "s3://nep-s3-xxxx/geospecies.rdf.gz",
"runNumber" : 1,
"retryNumber" : 0,
"status" : "LOAD_IN_PROGRESS",
"totalTimeSpent" : 148,
"startTime" : 1580199498,
"totalRecords" : 2130000,
"totalDuplicates" : 0,
"parsingErrors" : 0,
"datatypeMismatchErrors" : 0,
"insertErrors" : 0
}
}
}
Load Complete Output
[ec2-user@bastin nep-tool]$ curl -G 'https://neptest.xxxxxxxxxxxx.ap-northeast-1.neptune.amazonaws.com:8182/loader/eff1268f-17ab-473a-b845-c2d91a317c01'
{
"status" : "200 OK",
"payload" : {
"feedCount" : [
{
"LOAD_COMPLETED" : 1
}
],
"overallStatus" : {
"fullUri" : "s3://nep-s3-xxxx/geospecies.rdf.gz",
"runNumber" : 1,
"retryNumber" : 0,
"status" : "LOAD_COMPLETED",
"totalTimeSpent" : 149,
"startTime" : 1580199498,
"totalRecords" : 2201532,
"totalDuplicates" : 0,
"parsingErrors" : 0,
"datatypeMismatchErrors" : 0,
"insertErrors" : 0
}
}
The field descriptions are as follows. In this example, 2,201,532 records were loaded in 149 seconds.
Neptune Loader Get-Status API - Amazon Neptune https://docs.aws.amazon.com/ja_jp/neptune/latest/userguide/load-api-reference-status.html
| Field | Description |
|---|---|
| fullUri | The URI of one or more files to be loaded. Format: s3://bucket/key |
| runNumber | The number of runs for this load or feed. This increments when the load is resumed. |
| retryNumber | The number of retries for this load or feed. This increments when the loader automatically retries a feed or load. |
| status | The returned status for this load or feed. LOAD_COMPLETED indicates the load succeeded without issues. |
| totalTimeSpent | Time spent (in seconds) parsing and inserting data for this load or feed. Does not include time spent retrieving the list of source files. |
| totalRecords | Total records loaded or attempted to be loaded. |
| totalDuplicates | Number of duplicate records encountered. |
| parsingErrors | Number of parsing errors encountered. |
| datatypeMismatchErrors | Number of records where the specified data did not match the data type. |
| insertErrors | Number of records that could not be inserted due to errors. |
Issuing Queries to Neptune
Now that data is loaded, let’s issue queries.
Using the HTTP REST Endpoint
Connecting to a Neptune DB Instance Using the HTTP REST Endpoint - Amazon Neptune https://docs.aws.amazon.com/ja_jp/neptune/latest/userguide/access-graph-sparql-http-rest.html
curl -X POST --data-binary 'query=select ?s ?p ?o where {?s ?p ?o} limit 10' https://neptest.xxxxxxxxxxxx.ap-northeast-1.neptune.amazonaws.com:8182/sparql
Execution Result
[ec2-user@bastin nep-tool]$ curl -X POST --data-binary 'query=select ?s ?p ?o where {?s ?p ?o} limit 10' https://neptest.xxxxxxxxxxxx.ap-northeast-1.neptune.amazonaws.com:8182/sparql
{
"head" : {
"vars" : [ "s", "p", "o" ]
},
"results" : {
"bindings" : [ {
"s" : {
"type" : "uri",
"value" : "http://lod.geospecies.org/ses/uRtpv"
},
"p" : {
"type" : "uri",
"value" : "http://rdf.geospecies.org/ont/geospecies#isUnexpectedIn"
},
"o" : {
"type" : "uri",
"value" : "http://sws.geonames.org/5001836/"
}
~omitted~
Using the RDF4J Console
Connecting to a Neptune DB Instance Using the RDF4J Console - Amazon Neptune https://docs.aws.amazon.com/ja_jp/neptune/latest/userguide/access-graph-sparql-rdf4j-console.html
Download the RDF4J SDK from the RDF4J site .
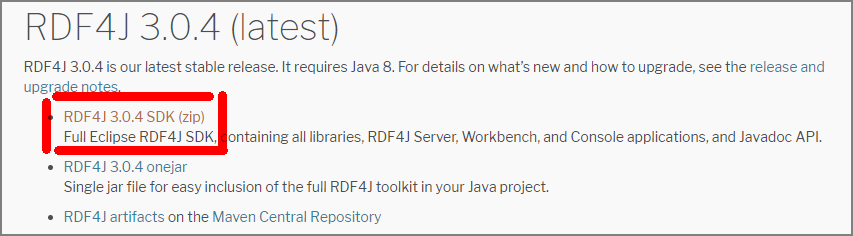
Upload the downloaded zip file to a specific EC2 instance.
[ec2-user@bastin nep-tool]$ ls -l
total 104740
-rw-r--r-- 1 ec2-user ec2-user 98147430 Jan 25 06:16 eclipse-rdf4j-3.0.4-sdk.zip
-rw-rw-r-- 1 ec2-user ec2-user 9105109 Jan 28 08:16 geospecies.rdf.gz
After unzipping, run console.sh located under bin/.
[ec2-user@bastin nep-tool]$ ./eclipse-rdf4j-3.0.4/bin/console.sh
08:37:35.639 [main] DEBUG org.eclipse.rdf4j.common.platform.PlatformFactory - os.name = linux
08:37:35.652 [main] DEBUG org.eclipse.rdf4j.common.platform.PlatformFactory - Detected Posix platform
Connected to default data directory
RDF4J Console 3.0.4+47737c0
3.0.4+47737c0
Type 'help' for help.
>
Create a SPARQL repository for the Neptune DB instance.
create sparql
You will be prompted to enter the following information. (Unverified, but if a read replica is created, the “SPARQL query endpoint” and “SPARQL update endpoint” should probably be split between master and read replica.)
| Variable | Value |
|---|---|
| SPARQL query endpoint | https://your-neptune-endpoint:port/sparql |
| SPARQL update endpoint | https://your-neptune-endpoint:port/sparql |
| Local repository ID [endpoint@localhost] | neptune |
| Repository title [SPARQL endpoint repository @localhost] | Neptune DB instance |
> create sparql
Please specify values for the following variables:
SPARQL query endpoint: https://neptest.xxxxxxxxxxxx.ap-northeast-1.neptune.amazonaws.com:8182/sparql
SPARQL update endpoint: https://neptest.xxxxxxxxxxxx.ap-northeast-1.neptune.amazonaws.com:8182/sparql
Local repository ID [endpoint@localhost]: neptune
Repository title [SPARQL endpoint repository @localhost]: neptune
Repository created
Connect to the Neptune instance. After connecting, the local repository ID appears in the prompt.
> open neptune
Opened repository 'neptune'
neptune>
Run the same query as with the HTTP REST endpoint approach.
neptune> sparql select ?s ?p ?o where {?s ?p ?o} limit 10
Evaluating SPARQL query...
+------------------------+------------------------+------------------------+
| s | p | o |
+------------------------+------------------------+------------------------+
| <http://lod.geospecies.org/ses/zJIK4>| <http://rdf.geospecies.org/ont/geospecies#hasScientificNameAuthorship>| "(LeConte, 1866)"^^xsd:string|
| <http://lod.geospecies.org/ses/zJIK4>| <http://rdf.geospecies.org/ont/geospecies#hasScientificName>| "Iphthiminus opacus (LeConte, 1866)"^^xsd:string|
| <http://lod.geospecies.org/ses/zJIK4>| <http://rdf.geospecies.org/ont/geospecies#isExpectedIn>| <http://sws.geonames.org/6255149/>|
| <http://lod.geospecies.org/ses/zJIK4>| <http://rdf.geospecies.org/ont/geospecies#isExpectedIn>| <http://sws.geonames.org/5279468/>|
| <http://lod.geospecies.org/ses/zJIK4>| <http://rdf.geospecies.org/ont/geospecies#hasNomenclaturalCode>| <http://rdf.geospecies.org/ont/geospecies#NomenclaturalCode_ICZN>|
| <http://lod.geospecies.org/ses/zJIK4>| <http://rdf.geospecies.org/ont/geospecies#isUnknownAboutIn>| <http://sws.geonames.org/4862182/>|
| <http://lod.geospecies.org/ses/zJIK4>| <http://rdf.geospecies.org/ont/geospecies#isUnknownAboutIn>| <http://sws.geonames.org/5037779/>|
| <http://lod.geospecies.org/ses/zJIK4>| <http://rdf.geospecies.org/ont/geospecies#isUnknownAboutIn>| <http://sws.geonames.org/5001836/>|
| <http://lod.geospecies.org/ses/zJIK4>| <http://rdf.geospecies.org/ont/geospecies#isUnknownAboutIn>| <http://sws.geonames.org/2635167/>|
| <http://lod.geospecies.org/ses/zJIK4>| <http://rdf.geospecies.org/ont/geospecies#hasSubfamilyName>| "Coelometopinae"^^xsd:string|
+------------------------+------------------------+------------------------+
10 result(s) (1268 ms)
neptune>